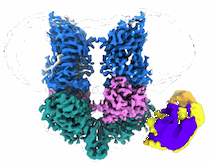Documentation
Summary
- EMDB data model
- EMDB header data model
- EMDB segmentation data model
- Policies
- Search engine
- Chart builder
- FAQ
- Deposition
EMDB map data model
The EM Data Bank (EMDB) accepts and distributes 3D map volumes derived from several types of EM reconstruction methods, including single particle averaging, helical averaging, 2D crystallography, and tomography. Since its inception in 2002, the EMDB map distribution format has followed CCP4 definition (CCP4 map format) , which is widely recognized by software packages used by the structural biology community. CCP4 map format is closely related to the MRC map format used in the 3DEM community (MRC map format); CCP4 is slightly more restrictive, in that voxel positions are limited to a grid that includes the Cartesian coordinate origin (0,0,0). Further details can be found here.
EMDB header data model
Every EMDB entry has a header file containing meta data (e.g., sample, detector, microscope, image processing) describing the experiment. The header file is an XML file and the structure and content of the header file is described by a XSD data model. With a highly dynamic field such as cryo-EM there is a constant need to adapt and modify the schema to keep it up-to-date with the most recent developments. We consult extensively with the EM community regarding such issues and version the schema according to the policy described here.
Data model version 1.9
This has been a long-term stable version of the data model. It was be replaced in 2018 with an updated model but XML header files in version 1.9 continues to be distributed in parallel for at least one year to give EMDB users ample time to switch. It should be noted that the generation of the version 1.9 header files will be on a best effort basis but involves a back translation from recent versions that are richer in content and will therefore not contain all the information that can be found in the more recent versions.
Download schema
Browse schema documentation
Download Python code to facilitate reading and writing XML version 1.9 header files
Data model version 3.0 (current model)
This data model replaced version 1.9, however header files corresponding to both data models will be distributed in parallel with the view of stopping the distribution of the version 1.9 files in 2019 once users have had a chance to adopt version 3.0.
This version adds a number of features including:
- An improved description of direct electron detectors, specimen preparation and tomography experiments.
- A hierarchal description of the overall sample composition in combination with a low-level description of the macromolecular composition to allow the description of both molecular and cellular samples.
- Specific data items describing the half-maps and segmentations included with the entry.
Download schema
Browse schema documentation
Download Python code to facilitate reading and writing XML version 1.9 header files
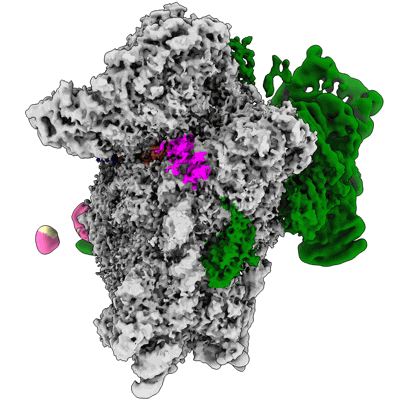
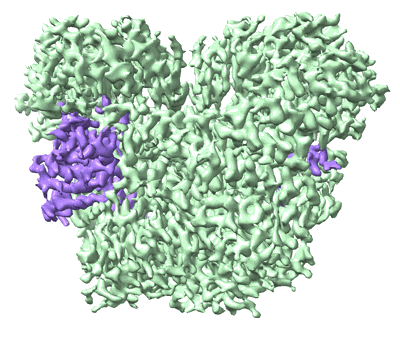
EMDB segmentation data model
Segmentation is the decomposition of 3D volumes into regions that can be associated with defined objects. Following several consultations with the EM community (Patwardhan et al., 2012; Patwardhan et al., 2014; Patwardhan et al., 2017), the EMDB is in the process of developing tools to support deposition of volume segmentations with structured biological annotation which is here defined as the association of data with identifiers (e.g., accession codes from UniProt) and ontologies taken from well established bioinformatics resources. To our knowledge, none of the segmentation formats widely used in electron microscopy and related fields currently support structured biological annotation. Third party use of segmentations is further impeded by the prevalence of segmentation file formats and their lack of interoperability. EMDB therefore proposed an open segmentation file format called EMDB-SFF to capture basic segmentation data from application-specific segmentation file formats and provide the means for structured biological annotation. In this way, EMDB-SFF will not only enable depositions of segmentations but also act as a file interchange format between different applications and facilitate analysis of 3D reconstructions. Furthermore EMDB-SFF supports the description of multiple transforms for a segment, thus allowing a segment to be used to describe the placement of a sub-tomogram average onto a tomographic reconstruction.
Model
EMDB-SFF files have the follow features:
- Segmentation metadata:
- name
- version (of schema)
- details (free-form text)
- global external references, e.g. specimen scientific identifier
- bounding box
- primary descriptor contained i.e. one of ‘three_d_volume’, ‘mesh_list’, or ‘shape_primitive_list’ (see schema documentation)
- list of software used to create the segmentation (name, version, processing details)
- list of transforms referenced by segments e.g. transform to place the sub-tomogram average in the tomogram
- Hierarchical ordering of segments through the use of segment IDs and parent IDs;
- Four geometrical representations of segments (volumes, contours, meshes, shapes);
- Can store subtomogram averages and how they map into the parent tomogram through the use of transforms;
- List of associated external references per segment;
- List of associated complexes and macromolecules in a related EMDB entry
Each segment in a segmentation can consist of two types of descriptors:
- textual descriptors;
- geometric descriptors.
Textual descriptors consist of either free-form text or standardised terms. Standard terms should be provided from a [published] ontology or list of identifiers.
Geometric descriptors can take one or more of the following representations:
- ‘three_d_volume’ for 3D volumes;
- ‘mesh_list’ for lists of meshes each of which consists of a set of vertices and polygons;
- lists of shape primitives (ellipsoid, cuboid, cone, cylinder).
Documentation
Download
The current schema (version 0.8.0.dev1) is available here.
Documentation
Complete documentation of the schema is available here.
Auxiliary Tools
sfftk-rw
sfftk-rw is a Python toolkit for reading and writing EMDB-SFF files only. It is part of a family of tools designed to work with EMDB-SFF files.
sfftk-rw has the following utilities:
- convert - interconvert between XML, HDF5 and JSON file formats of the EMDB-SFF data model;
- view - view a file summary
The full documentation is available at readthedocs.
Download
The latest version runs only on Python 3 (version 0.7.1) and may be installed using pip install sfftk-rw. Alternatively, feel free to obtain the source code from Github.
sfftk
sfftk provides a shell command and a Python API to process EMDB-SFF files.
The following utilities are available using sfftk:
- convert - Conversion of application-specific segmentation file formats to EMDB-SFF. Currently, sfftk supports the following formats:
- AmiraMesh (.am)
- Amira HyperSurface (.surf)
- Segger (.seg)
- EMDB Map masks (.map)
- Stereolithography (.stl)
- IMOD (.mod)
- notes - Annotation of EMDB-SFF files.
- view - Brief summaries of segmentation files.
Read the full documentation here.
Download
The latest development version (version 0.5.5.dev1) of sfftk may be downloaded/installed from PyPI or the source may be obtained from GitHub.
Publications
- Patwardhan, Ardan, Robert Brandt, Sarah J. Butcher, Lucy Collinson, David Gault, Kay Grünewald, Corey Hecksel et al. Building bridges between cellular and molecular structural biology. eLife 6 (2017).
- Patwardhan, Ardan, Alun Ashton, Robert Brandt, Sarah Butcher, Raffaella Carzaniga, Wah Chiu, Lucy Collinson et al. A 3D cellular context for the macromolecular world. Nature structural & molecular biology 21, no. 10 (2014): 841-845.
- Patwardhan, Ardan, José-Maria Carazo, Bridget Carragher, Richard Henderson, J. Bernard Heymann, Emma Hill, Grant J. Jensen et al. Data management challenges in three-dimensional EM. Nature structural & molecular biology 19, no. 12 (2012): 1203-1207.
Quick links
Recent Entries
(Show all)Cryo-EM of the GDP-bound human dynamin polymer assembled on the membrane in the super constricted state
Cryo-EM structure of human exon-defined spliceosome in the late pre-B state.
Cryo-EM structure of SV2A LD4 in complex with BoNT/A2 Hc in the SV2A-levetiracetam-BoNT/A2 Hc complex
Structure of the human neutral amino acid transporter ASCT2 in complex with nanobody 469
Endogenous trans-translation complex with tmRNA*SmpB in the P site and alanyl-tRNA in the A site of E. coli 70S ribosome
TUBB4B and TUBA1A Heterodimer from Human Respiratory Doublet Microtubules
Cryo-EM structure of Mycobacterium tuberculosis ATP synthase Fo in complex with bedaquiline(BDQ)
Roco protein from C. tepidum in the GTP state bound to the activating Nanobodies NbRoco1 and NbRoco2
60S ribosome biogenesis intermediate (Dbp10 catalytic structure - Dbp10 Local map)
60S ribosome biogenesis intermediate (Dbp10 catalytic structure - L1 local map
Open State of central tail fiber of bacteriophage lambda upon binding to LamB (gpJ713-LamB complex)
Staphylococcus aureus 70S ribosome with elongation factor G locked with fusidic acid cyclopentane with a tRNA in pe/E chimeric hybrid state
Staphylococcus aureus 70S ribosome with elongation factor G locked with fusidic acid with a tRNA in pe/E chimeric state
Cryo-EM structure of human exon-defined spliceosome in the early B state.
60S ribosome biogenesis intermediate (Dbp10 post-catalytic structure - Dbp10 Local map)
Cryo-EM Structure of Spike Glycoprotein from Civet Coronavirus 007 in Closed Conformation
Cryo-EM structure of the Pseudomonas aeruginosa PAO1 Type IV pilus
Cryo-EM of the GDP-bound human dynamin (full-length) polymer assembled on the membrane in the super constricted state (full helix)
Cryo-EM of the GDP-bound human dynamin polymer assembled on the membrane in the super constricted state showing the PH domain
Structure of the human mitochondrial iron-sulfur cluster biosynthesis complex during persulfide transfer (consensus map)
60S ribosome biogenesis intermediate (Dbp10 pre-catalytic structure - Overall map)
Human DNA polymerase theta helicase domain tetramer in the apo form
Structure of the human mitochondrial iron-sulfur cluster biosynthesis complex during persulfide transfer (without frataxin)
Cryo-EM structure of FLVCR2 in the inward-facing state with choline bound
Cryo-EM structure of SV2A in complex with BoNT/A2 Hc and levetiracetam
Chlorella virus Hyaluronan Synthase bound to GlcA extended GlcNAc primer and UDP
Staphylococcus aureus 70S ribosome with elongation factor G locked with fusidic acid cyclopentane in post-translocational state
2up-TM conformation of HKU1-B S protein after incubation of the receptor
Structure of the human ATP synthase bound to bedaquiline (peripheral stalk domain)
Structure of the human ATP synthase bound to bedaquiline (composite)
Mycobacterium tuberculosis ATP synthase Peripheral Stalk in the apo-form
60S ribosome biogenesis intermediate (Dbp10 post-catalytic structure - H64 Local map)
Cryo-EM structure of FLVCR2 in the outward-facing state with choline bound
2up-1 conformation of HKU1-B S protein after incubation of the receptor
Cryo-EM Structure of CdnG-E2 complex from Serratia marcescens (UltrAuFoil)
60S ribosome biogenesis intermediate (Dbp10 pre-catalytic structure - Local map Rrp14/Rrp15/Ssf1 region)
Cryo-EM structure of SV2A dimer in complex with BoNT/A2 Hc and levetiracetam
3up-TM conformation of HKU1-B S protein after incubation of the receptor
Cryo-EM structure of conformation 1 of complex of Nipah virus attachment glycoprotein G with 1E5 neutralizing antibody
Cryo-EM Structure of Spike Glycoprotein from Bat Coronavirus WIV1 in Closed Conformation
Structure of Xenopus tropicalis acid-sensitive outwardly rectifying channel ASOR trimer bound with tRNA (intermediate state)
60S ribosome biogenesis intermediate (Dbp10 catalytic intermediate - Rrp14/Rrp15/Ssf1 local map)
Closed conformation of HKU1-B S protein after incubation of the receptor
1up-2 conformation of HKU1-B S protein after incubation of the receptor
Xenopus laevis hyaluronan synthase 1, UDP-bound, gating loop inserted state
Chlorella virus Hyaluronan Synthase bound to GlcNAc primer and UDP-GlcA
Structure of the human mitochondrial iron-sulfur cluster biosynthesis complex during persulfide transfer (persulfide on NFS1 and ISCU2)
The Anoxybacillus pushchinoensis ORF-less Group IIC Intron HYER1 with 10-nt TRS at symmetric apo state
Xenopus laevis hyaluronan synthase 1, nascent HA polymer bound state
Cryo-EM structure of Mycobacterium tuberculosis ATP synthase Fo in the apo-form
60S ribosome biogenesis intermediate (Dbp10 catalytic structure - Overall map)
Ensemble map of the Roco protein from C. tepidum in the GTP state bound to the activating Nanobodies NbRoco1 and NbRoco2
CRYO-EM FOCUSED REFINEMENT MAPS OF LEISHMANIA MAJOR 80S RIBOSOME : WILD TYPE REPLICATE 1
Chlorella virus Hyaluronan Synthase bound to GlcA extended GlcNAc primer
Open state of central tail fiber of bacteriophage lambda upon binding to LamB
Roco protein from C. tepidum in the GTP state bound to an activating Nanobody
Cryo-EM Structure of Smooth Muscle Gamma Actin (ACTG2) Mutant R257C
60S ribosome biogenesis intermediate (Dbp10 post-catalytic structure - Overall map)
Cryo-EM structure of a Legionella effector complexed with actin and AMP
type I-B Cascade bound to a PAM-containing dsDNA target at 3.8 angstrom resolution.
60S ribosome biogenesis intermediate (Dbp10 pre-catalytic structure - Local map L1 region)
Human DNA polymerase theta helicase domain dimer bound to DNA in the microhomology aligning conformation
Mycobacterium tuberculosis ATP synthase F1 in complex with bedaquiline(BDQ)
Human DNA polymerase theta helicase domain dimer bound to DNA in the microhomology searching conformation
Structure of the dimeric Xenopus tropical acid-sensitive outwardly rectifying channel ASOR trimer bound with tRNA (closed state)
Cryo-EM structure of Mycobacterium tuberculosis ATP synthase F1 in complex with TBAJ-587
Serotonin 1E receptor (5-HT1eR)-Gi1 Complex bound with Setiptiline
Cryo-EM structure of a bacterial nitrilase filament with a covalent adduct derived from benzonitrile hydrolysis
Cryo-EM structure of SV2A in complex with BoNT/A2 Hc and brivaracetam
60S ribosome biogenesis intermediate (Dbp10 catalytic structure - Low-pass filtered locally refined map)
Structure of the human mitochondrial iron-sulfur cluster biosynthesis complex during persulfide transfer (persulfide on ISCU2)
Cryo-EM of the GDP-bound human dynamin polymer assembled on the membrane in the super constricted state (full helix)
Cryo-EM structure of a Legionella effector complexed with actin and ATP
1up-1 conformation of HKU1-B S protein after incubation of the receptor
Structure of the human ATP synthase bound to bedaquiline (membrane domain)
Microtubule inner proteins in the 48-nm doublet microtubule from the proximal region of Tetrahymena thermophila strain K40R
Cryo-EM structure of the the 2-oxoglutarate dehydrogenase (E1) with TCAIM complex
Cryo-EM structure of human exon-defined spliceosome in the mature pre-B state.
cryoEM map for design HE0537, a D4 symmetric homo-oligomer designed with RFdiffusion.
Structure of the SARS-CoV-2 EG.5.1 spike glycoprotein in complex with ACE2 (1-up state)
Cryo-EM structure of conformation 2 of complex of Nipah virus attachment G with 1E5 neutralizing antibody
Cryo-EM structure of Mycobacterium tuberculosis ATP synthase Peripheral Stalk in complex with TBAJ-587
60S ribosome biogenesis intermediate (Dbp10 pre-catalytic structure - PTC Local map)
Cryo-EM of the GDP-bound human dynamin (full-length) polymer assembled on the membrane in the super constricted state
Structure of Xenopus tropicalis acid-sensitive outwardly rectifying channel ASOR (resting state)
Cryo-EM of the GDP-bound human dynamin polymer assembled on the membrane in the super constricted state showing the second PH domain
Endogenous trans-translation complex with tmRNA*SmpB in the P site and alanyl-tRNA in the A site and deacyl-tRNA in the E site of E. coli 70S ribosome
Monkeypox virus DNA replication holoenzyme F8, A22 and E4 complex in a DNA binding form
Mycobacterium tuberculosis ATP synthase Peripheral Stalk in complex with bedaquiline(BDQ)
Focused map on the Roc-COR domains of the Roco protein from C. tepidum in the GTP state bound to the activating Nanobody NbRoco1
Cryo-EM structure of the gasdermin pore from Trichoplax adhaerens
Cryo-EM structure of Mycobacterium tuberculosis ATP synthase Fo in complex with TBAJ-587
Cryo-EM structure of Mycobacterium tuberculosis ATP synthase in complex with TBAJ-587
Structural mechanism of inhibition of the Rho transcription termination factor by Rof
Human DNA polymerase theta helicase domain dimer bound to DNA in the microhomology annealed conformation
Cryo-EM structure of human exon-defined spliceosome in the mature B state.
Focused map on the LRR domain of the Roco protein from C. tepidum bound to the activating Nanobody NbRoco2
Cryo-EM structure of SV2A in complex with BoNT/A2 Hc and levetiracetam